Maximum likelihood
This post is one of those ‘explain to myself how things work’ documents, which are not necessarily completely correct but are close enough to facilitate understanding.
Background #
Let’s assume that we are working with a fairly simple linear model, where we only have a response variable (say tree stem diameter in cm). If we want to ‘guess’ the diameter for a tree (yi) our best bet is the average (μ) and we will have a residual (εi). The model equation then looks like:
\(y_i = \mu + \varepsilon_i\)
We want to estimate both the overall mean (\(\mu\)) and the error variance (\(\sigma_{\varepsilon}^2\)), for which we could use least squares. However, we will use an alternative method (maximum likelihood) because that is the point of this post. A likelihood function expresses the probability of obtaining the observed sample from a population given a set of model parameters. Thus, it answers the question What is the probability of observing the current dataset, when I assume a given set of parameters for my linear model? To do so, we have to assume a distribution, say normal, for which the probability density function (p.d.f) is:
\(p.d.f. (y) = \frac{1}{\sqrt{2 \pi} \sigma} e^{-1/2 \frac{(y- \mu)^2}{\sigma^2}}\)
with n independent samples the likelihood (L) is:
\(L = \prod_{i=1}^n \frac{1}{\sqrt{2 \pi} \sigma} e^{-1/2 \frac{(y- \mu)^2}{\sigma^2}}\)
where \(\prod\) is a multiplication operator, analogous to the summation operator \(\sum\). Maximizing L or its natural logarithm (LogL) is equivalent, then:
$$\log L = \sum_{i=1}^n \log \lparen \frac{1}{\sqrt{2 \pi} \sigma} e^{-1/2 \frac{(y- \mu)^2}{\sigma^2}} \rparen \\ \log L= \sum_{i=1}^n \lbrack \log \lparen \frac{1}{\sqrt{2 \pi} \sigma} \rparen + \log \lparen e^{-1/2 \frac{(y- \mu)^2}{\sigma^2}} \rparen \rbrack \\ \log L= \sum_{i=1}^n \log(2 \pi)^{-1/2} + \sum_{i=1}^n \log \lparen \frac{1}{\sigma} \rparen + \sum_{i=1}^n \lparen -1/2 \frac{(y- \mu)^2} {\sigma^2} \rparen \\ \log L= -\frac{n}{2} \log(2 \pi) – n \log(\sigma) – \frac{1}{2 \sigma^2} \sum_{i=1}^n ( y – \mu)^2)$$Considering only \(\mu\), \(\log L\) is maximum when \(\sum_{i=1}^n ( y – \mu)^2\) is a minimum, i.e. for:
$$\mu = \frac{\sum_{i=1}^n y_i}{n} $$Now considering \(\sigma \):
\(\frac{\delta LogL}{\delta \sigma} = -\frac{n}{\sigma} – (-2) \frac{1}{2} \sum_{i=1}^n (y – \mu)^2 \sigma^{-3}\)
\(\frac{\delta LogL}{\delta \sigma} = -\frac{n}{\sigma} + \frac{1}{\sigma^3} \sum_{i=1}^n (y – \mu)^2\)
and setting the last expression equal to 0:
\(\frac{n}{\sigma} = \frac{1}{\sigma^3} \sum_{i=1}^n (y – \mu)^2\)
\(\sigma^2 = \frac{\sum_{i=1}^n (y – \mu)^2}{n}\)
We can obtain an estimate of the error variance (\(\hat{\sigma}\), notice the hat) by replacing \(\mu\) by our previous estimate:
\(\hat{\sigma}^2 = \frac{sum_{i=1}^n (y – \bar{y})^2}{n}\)
which is biased, because the denominator is n rather than (n – 1) (the typical denominator for sample variance). This bias arises because maximum likelihood estimates do not take into account the loss of degrees of freedom when estimating fixed effects.
Playing in R with an example #
We have data for stem diameters (in mm) for twelve 10 year-old radiata pine (Pinus radiata D. Don) trees:
diams <- c(150, 124, 100, 107, 170, 144,
113, 108, 92, 129, 123, 118)
# Obtain typical mean and standard deviation
mean(diams)
[1] 123.1667
sd(diams)
[1] 22.38438
# A simple function to calculate likelihood.
# Values quickly become very small
like <- function(data, mu, sigma) {
like <- 1
for(obs in data){
like <- like * 1/(sqrt(2*pi)*sigma) * exp(-1/2 * (obs - mu)^2/(sigma^2))
}
return(like)
}
# For example, likelihood of data if mu=120 and sigma=20
like(diams, 120, 20)
[1] 3.476384e-24
# A simple function to calculate log-likelihood.
# Values will be easier to manage
loglike <- function(data, mu, sigma) {
loglike <- 0
for(obs in data){
loglike <- loglike + log(1/(sqrt(2*pi)*sigma) * exp(-1/2 * (obs - mu)^2/(sigma^2)))
}
return(loglike)
}
# Example, log-likelihood of data if mu=120 and sigma=20
loglike(diams, 122, 20)
[1] -53.88605
# Let's try some combinations of parameters and
# plot the results
params <- expand.grid(mu = seq(50, 200, 1),
sigma <- seq(10, 40, 1))
params$logL <- with(params, loglike(diams, mu, sigma))
summary(params)
library(lattice)
contourplot(logL ~ mu*sigma, data = params, cuts = 20)
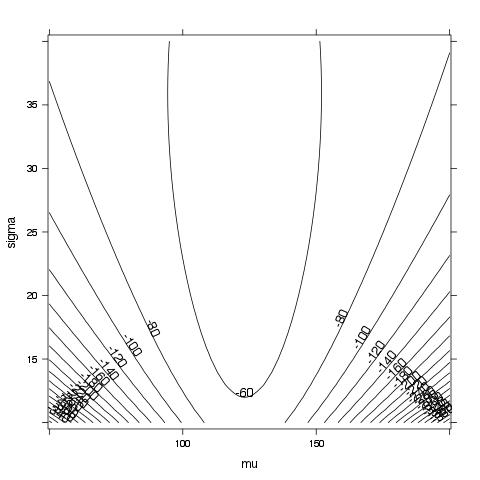
It is difficult to see the maximum likelihood in this plot, so we will zoom-in by generating a smaller grid around the typical mean and standard deviation estimates.
Zooming in #

We can now check what would be actual results using any of the functions to fit models using maximum likelihood, for example gls():
library(nlme)
m1 <- gls(diams ~ 1, method='ML')
summary(m1)
Generalized least squares fit by maximum likelihood
Model: diams ~ 1
Data: NULL
AIC BIC logLik
111.6111 112.5809 -53.80556
Coefficients:
Value Std.Error t-value p-value
(Intercept) 123.1667 6.461815 19.06069 0
Residual standard error: 21.43142
Degrees of freedom: 12 total; 11 residual
In real life, software will use an iterative process to find the combination of parameters that maximizes the log-likelihood value. If we go back to our function and use loglike(diams, 123.1667, 21.43142) we will obtain -53.80556; exactly the same value calculated by gls. In addition, we can see that the estimated standard deviation (21.43) is slightly lower than the one calculated by the function sd() (22.38), because our biased estimate divides the sum of squared deviations by n rather than by n-1.
P.S. The code for the likelihood and log-likelihood functions is far from being optimal (the loops could be vectorized). However, the loops are easier to understand for many people.